Libraries preparation
A complete of 265 medicinal vegetation had been chosen and launched into the ChEBI database to extract the efficient compounds. After eradicating duplicates, 87 potential compounds had been obtained from the 265 native plant species. The PharmMapper web site has launched 423 potential goal candidates for these compounds. As well as, the NCBI database has launched 3312 colorectal cancer-related targets. Lastly, 34 human protein molecules had been chosen, which may very well be targets of those compounds and play a task in colorectal most cancers.
Of those compounds, 7 didn’t have widespread targets of their high 30 targets. The names and 2D buildings of each the teams of compounds used within the screening and their targets are listed in Supplementary Tables S1âS3.
DT community development and evaluation
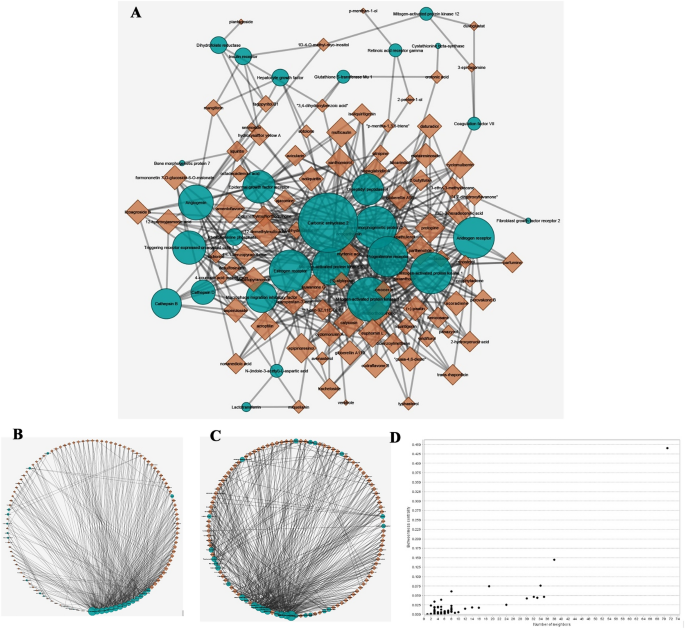
A DT community with 87 compounds and 34 targets was constructed consisting of 114 nodes and 416 levels. The NetworkAnalyser outcomes calculate parameters such because the variety of levels, betweenness, and edge betweenness centrality values, that are used to pick one of the best molecular targets and potential chemical compounds (Fig. 1). Figure 1A exhibits a bipartite community configured with node sizes primarily based on the variety of levels; the bigger the node dimension, the larger is the variety of associations. Based mostly on the community evaluation, the compounds confirmed levels starting from 1 to eight. To display one of the best compounds as ligands for molecular docking evaluation in opposition to targets, compounds with six or extra nodes had been chosen. Due to this fact, 31 compounds listed in Desk 1 had been chosen for additional examine. Carbonic anhydrase 2, with a rating of 0.44, confirmed the best centrality among the many targets, adopted by mitogen-activated protein kinase 14 (0.14), estrogen receptor (0.07), and angiogenin (0.07). The remaining molecules had a rating of lower than 0.05. Furthermore, Carbonic anhydrase 2 (71), mitogen-activated protein kinase 14 (38), bone morphogenetic protein 2 (35), and estrogen receptor (34) confirmed essentially the most connections, primarily based on the variety of levels related to the nodes.
(A) The bipartite DT community. (B) The circle structure of DT community primarily based on diploma. (C) The circle structure of DT community primarily based on betweenness centrality. (D) The betweenness centrality of nods. The inexperienced round nodes symbolize the targets, and the brown triangular nodes symbolize the chemical compounds. The bigger the scale of the node, the larger the variety of nodes and associations.
Lastly, the compounds confirmed numerous levels starting from 1 to eight. To display one of the best compounds as ligands for the molecular docking step in opposition to the targets, all compounds with six or extra nodes had been chosen, wherein there have been 31 compounds with six, seven, or eight nodes.
Molecular docking and simulation
After checking utilizing the GEPIA database, the TOP targets that confirmed elevated expression in colorectal most cancers had been chosen. The goal buildings had been downloaded from the PDB database (PDB IDs: 6AY2 for CTSB, 5T4E for DPP4, 6FVE for MIF, 7AUV for MAPK1, 4QTD for MAPK8, and 1SMO for TERM1). The Chimera 1.8.1 software program was used for important protein preparations, together with eradicating water, ATP, ligands, and including hydrogen costs. Essential ligand-binding websites had been thought-about and docked to 31 compounds utilizing the PyRx software program. 31 molecules had been chosen after analyzing the pharmacokinetic properties and parameters of ADME. The centroid of the binding websites for the targets was calculated because the coordinates of the centroid of the ligand-binding websites utilizing the UniProt database. The docked complexes had been analyzed utilizing PyMol and DiscoveryStudio software program. Lastly, molecules that offered the best interplay vitality in opposition to their goal (above ââ7.5 for TERM1, ââ8 for CTSB and MIF, above ââ9 for DPP4 and MAPK1 and above ââ10 for MAPK8), respectively and had extra binding with the amino acids of the binding website within the examine with DiscoveryStudio. These findings led to the choice of Multiorthoquinone, Liquiritin, Hispaglabridin A, Isoliquiritin, Gibberellin A98, Cyclomulberrin, Cyclomorusin A, Cudraflavone B for simulation (MD), which produced a extra secure advanced with a decrease vitality stage than TREM1, MAPK8, MAPK1, MAPK8, CTSB, MAPK1, MIF, and DPP4, respectively. The positions and amino acids concerned within the binding are illustrated in Fig. 2, 3, 4, 5, 6, 7, 8 and 9, Supplementary Tables S4âS6. The RMSD rating confirmed slight fluctuations and was roughly 0.2 nm. These findings point out the steadiness of the compounds within the goal advanced. The RMSF values for all protein buildings had been computed to precisely decide how the binding of the compounds impacts flexibility.
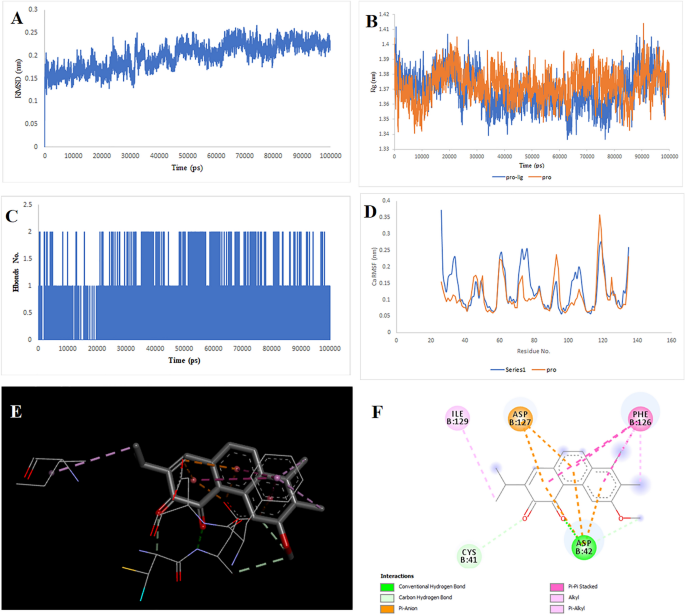
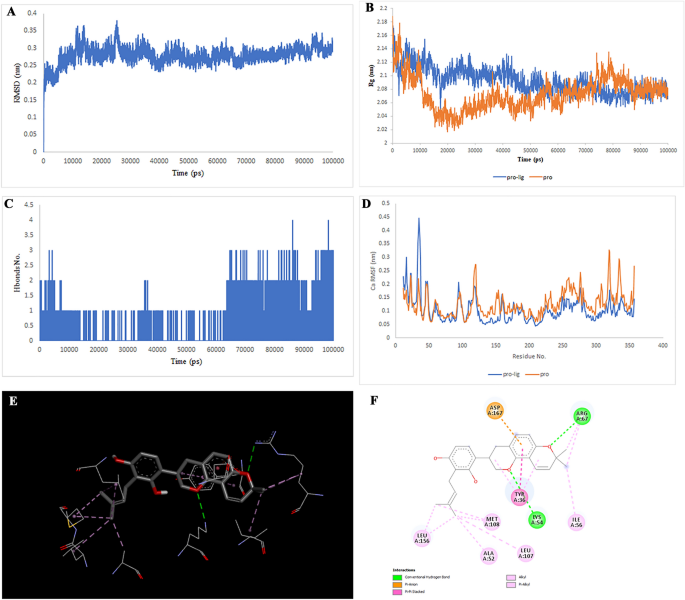
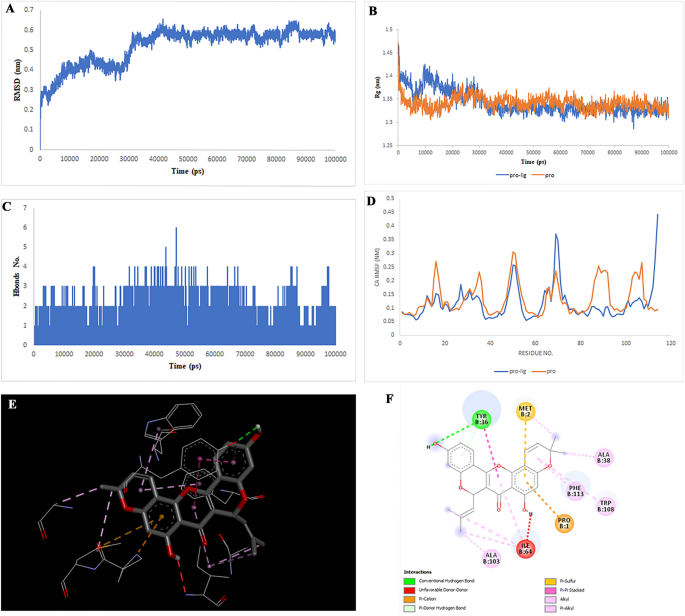
Two-dimensional representations of the Multiorthoquinone in opposition to TREM1. (A) Root-mean sq. deviation of the complexes (RMSD). (B) Radius of gyration (Rg). (C) Hydrogen bond evaluation from the simulation system. (D) Root-mean-square fluctuation (RMSF). (E) The binding conformation of 3D view. (F) Binding website interactions of 2D view.
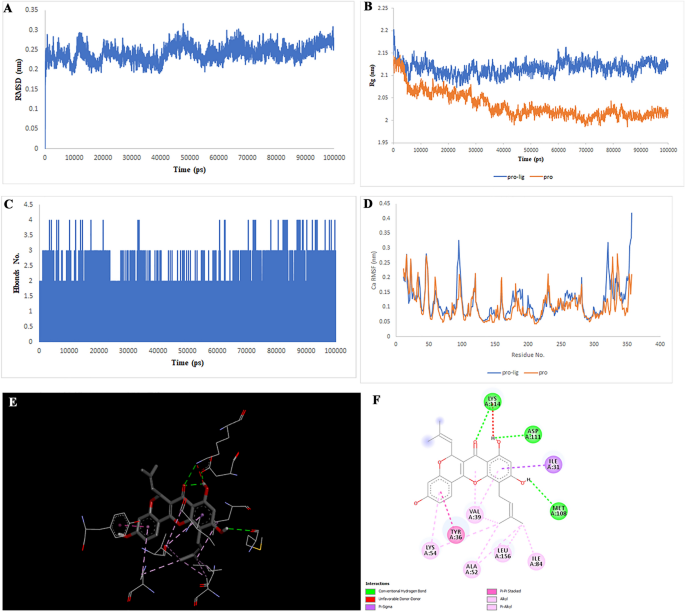
Two-dimensional representations of the Liquiritin in opposition to MAPK8. (A) Root-mean sq. deviation of the complexes (RMSD). (B) Radius of gyration (Rg). (C) Hydrogen bond evaluation from the simulation system. (D) Root-mean-square fluctuation (RMSF). (E) The binding conformation of 3D view. (F) Binding website interactions of 2D view.
Two-dimensional representations of the Isoliquiritin in opposition to MAPK8. (A) Root-mean sq. deviation of the complexes (RMSD). (B) Radius of gyration (Rg). (C) Hydrogen bond evaluation from the simulation system. (D) Root-mean-square fluctuation (RMSF). (E) The binding conformation of 3D view. (F) Binding website interactions of 2D view.
Two-dimensional representations of the Hispaglabridin A in opposition to MAPK1. (A) Root-mean sq. deviation of the complexes (RMSD). (B) Radius of gyration (Rg). (C) Hydrogen bond evaluation from the simulation system. (D) Root-mean-square fluctuation (RMSF). (E) The binding conformation of 3D view. (F) Binding website interactions of 2D view.
Two-dimensional representations of the Cyclomulberrin in opposition to MAPK1. (A) Root-mean sq. deviation of the complexes (RMSD). (B) Radius of gyration (Rg). (C) Hydrogen bond evaluation from the simulation system. (D) Root-mean-square fluctuation (RMSF). (E) The binding conformation of 3D view. (F) Binding website interactions of 2D view.
Two-dimensional representations of the Cudraflavone B in opposition to DPP4. (A) Root-mean sq. deviation of the complexes (RMSD). (B) Radius of gyration (Rg). (C) Hydrogen bond evaluation from the simulation system. (D) Root-mean-square fluctuation (RMSF). (E) The binding conformation of 3D view. (F) Binding website interactions of 2D view.
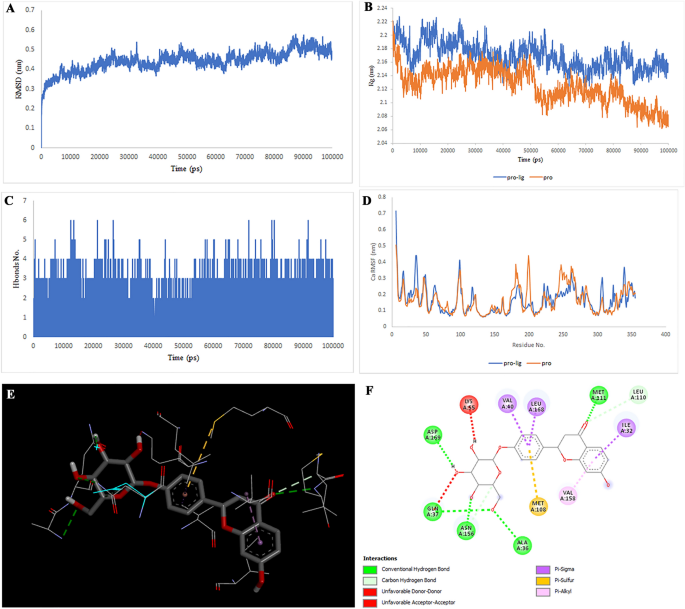
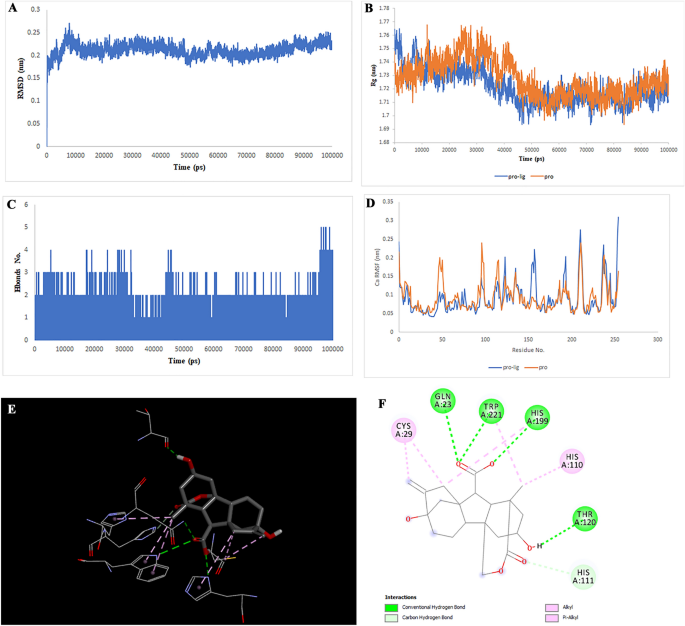
Two-dimensional representations of the Gibberellin A98 in opposition to CTSB. (A) Root-mean sq. deviation of the complexes (RMSD). (B) Radius of gyration (Rg). (C) Hydrogen bond evaluation from the simulation system. (D) Root-mean-square fluctuation (RMSF). (E) The binding conformation of 3D view. (F) Binding website interactions of 2D view.
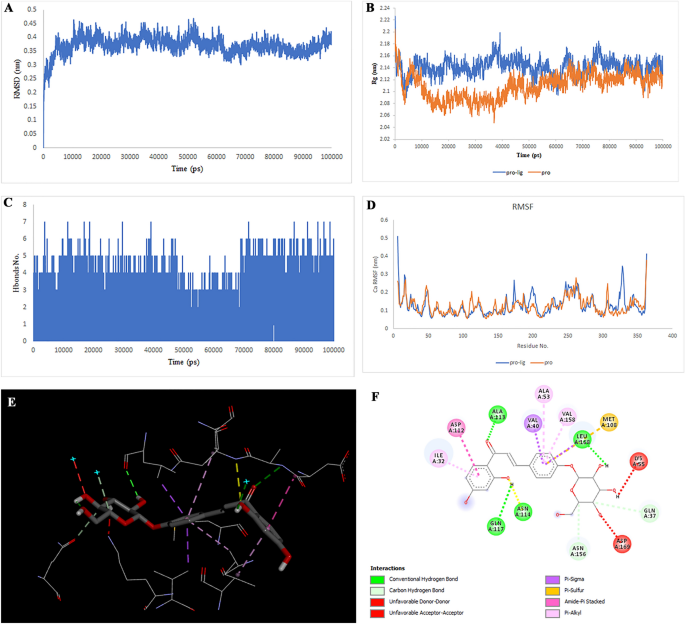
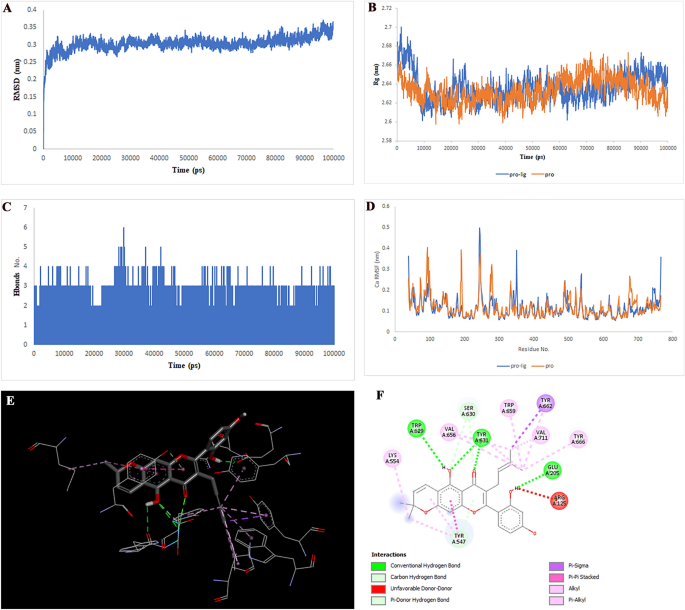
Two-dimensional representations of the Cyclomulberrin in opposition to MIF. (A) Root-mean sq. deviation of the complexes (RMSD). (B) Radius of gyration (Rg). (C) Hydrogen bond evaluation from the simulation system. (D) Root-mean-square fluctuation (RMSF). (E) The binding conformation of 3D view. (F) Binding website interactions of 2D view.
Evaluation of the liquiritin- and isoliquiritin-associated RMSF plots revealed that Liquiritin and Isoliquiritin flexibility had been considerably completely different within the 5 areas of 35, 183, 199, 246, and 307 and two areas in 200 and 328 in MAPK8, respectively. As well as, multiorthoquinone flexibility was extraordinarily excessive within the three areas 34, 76, and 107 in TREM1, which can be attributed to an absence of interplay between the three areas and multiorthoquinone. For Cyclomulberrin and Hispaglabridin A residues 94â320, and 35, 300â350 in MAPK1, the pliability of amino acids was decrease and better, respectively. The RMSF plot related to DPP4 indicated excessive interplay between most areas of DPP4 and Cudraflavone B. Moreover, Cyclomorusin A and Gibberellin A98 flexibility had been extraordinarily completely different within the two areas of 50â70 and 150â170 within the CTSB, and 80â100 area within the MIF, respectively.
RG nature was fixed for the person domains throughout the complete simulation interval related to Gibberellin A98, Cyclomorusin A, and Cudraflavone B, indicating that the person domains didn’t soften or unfold. These compounds didn’t have an effect on the secondary buildings of CTSB, MIF, or DPP4. Nevertheless, the RG worth related to Liquiritin, Isoliquiritin and Hispaglabridin A, Cyclomulberrin all through the MD simulation led to unfolding and activation of MAPK8 and MAPK1, respectively.
Principal element evaluation
Supplementary Fig. S1 exhibits the projection of the trajectories on particular eigenvectors (vectors 1 and a couple of) and time-dependent motions of the parts in a specific vibration mode. The general evaluation of the eigenvector plots indicated that the majority vibrations occurred alongside eigenvector 1. In line with the primary two PCs, TREM1 and MIF proteins have nearly the identical hint values of the covariance matrix for the certain and unbound states with a slight shift. This means that the ligand is properly equilibrated and stabilized with the protein, as mirrored by theleast conformational modifications attributable to decreased collective motions from unbound states. Nevertheless, in different circumstances, 2D projection plots of the trajectories revealed that the ligand decreased the conformational variety in the course of the simulations, resulting in a extra compact cluster distribution. Sampling of various areas and displaying completely different motion behaviors of proteinâligand complexes in comparison with unbound proteins factors to the binding of ligand results on the rigidity of the structural conformation, which additionally impacts the operate of proteins.
MM/PBSA binding free vitality
The MM/PBSA binding free vitality outcomes are proven in Desk 2 together with the van der Waals vitality (kJ/mol), electrostatic vitality (kJ/mol), polar solvation vitality (kJ/mol), SASA vitality (kJ/mol), SAV vitality (kJ/mol), WCA vitality (kJ/mol), and binding vitality (kJ/mol), are proven in Desk 2. In comparison with liquiritin, isoliquiritin had the bottom binding vitality rating of 183.04Â kJ/mol interacting with MAPK8 and fashioned a stronger binding. As well as, hypoglabrin A and cyclomulbrin work together with MAPK1 at nearly the identical binding vitality (ââ154Â kJ/mol). Nevertheless, the variety of hydrogen bonds within the interplay with hypoglabrin A is comparatively excessive. MIF-Cyclomorusin A, with essentially the most detrimental vitality rating (ââ243.768Â kJ/mol), confirmed the strongest interplay amongst all docked complexes.









Discussion about this post